About Me
Originally a math major, I trained with Dr. Calin Belta (BU) and Dr. Ron Weiss (MIT), earning my doctorate degree in bioinformatics from Boston University. After completing my postdoctoral research at University of Southern California, I joined AstraZeneca as a bioinformatics scientist to further my career in biomedical research. My primary research focuses on pipeline automation, application of statistical methods to omics data analysis, as well as machine learning and artificial intelligence to stratify patients, predict drug-target interactions, and assess drug toxicity.
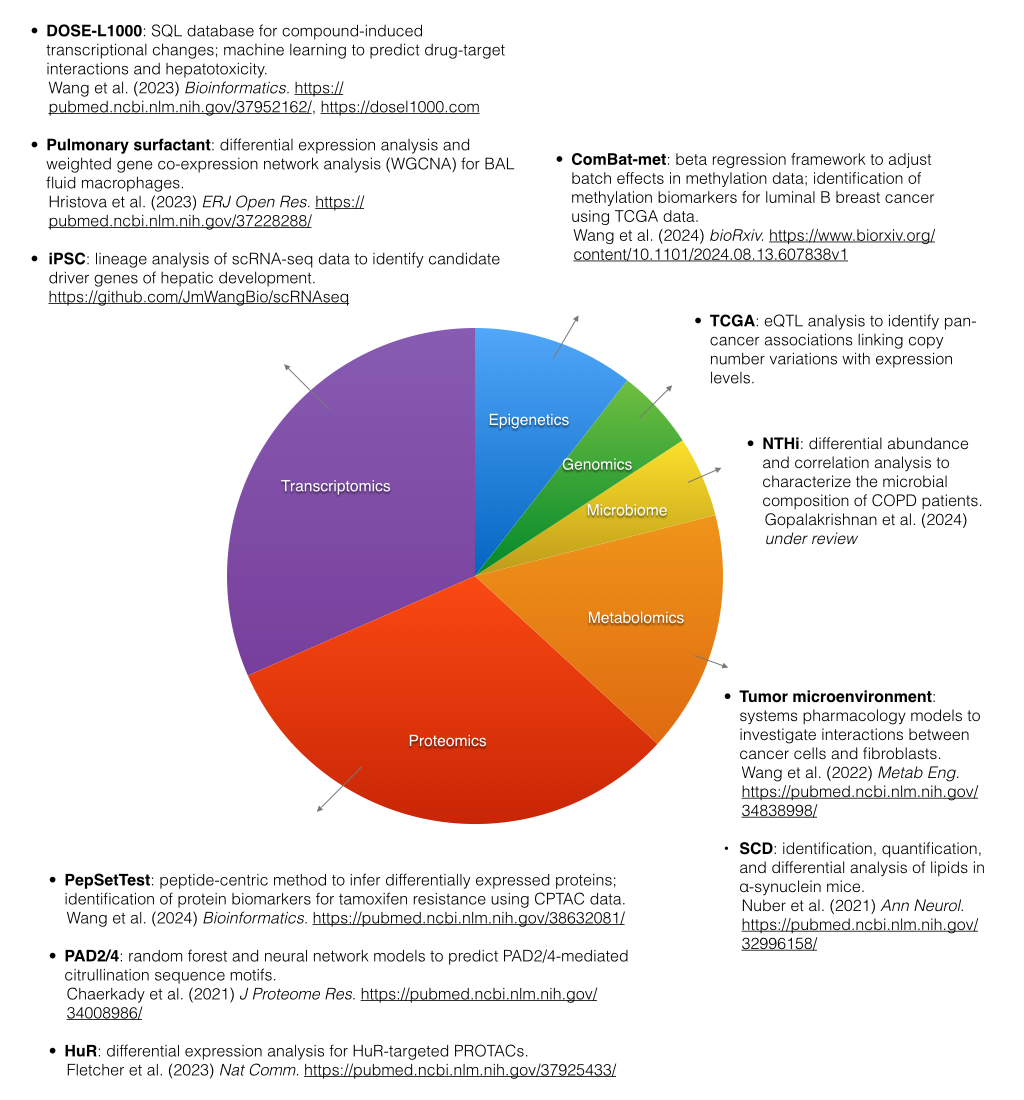
Project Highlights
1) Developed generalized additive models (a type of statistical learning) to calculate the estimates and standard errors of gene expression changes, efficacy, and potency across 33,395 compounds, 82 cell lines, and 978 genes.
2) Designed DOSE-L1000, an SQL-based database and DOSE-L1000-Viz, a Shiny-based application to visualize and analyze compound-induced transcriptional changes and dose-response data.
3) Leveraged random forests and neural networks to predict drug-target interactions and organ-specific drug toxicity from drug-induced expression changes.
- Keywords: Transcriptomics, Machine Learning
- GitHub: https://github.com/JmWangBio/DoseL1000
- Website: https://dosel1000.com
- Publication: Wang J., Novick S., “DOSE-L1000: Unveiling the Intricate Landscape of Compound-Induced Transcriptional Changes.” Bioinformatics, 2023, 39(11): btad683. doi: 10.1093/bioinformatics/btad683
1) Developed ComBat-met, a beta regression framework as an R package, to adjust batch effects in DNA methylation data.
2) Utilized neural networks to predict breast cancer subtypes from TCGA methylation data, following batch-effect adjustment using ComBat-met.
- Keywords: Epigenetics, Statistics, Software Development
- GitHub: https://github.com/JmWangBio/ComBatMet
- Preprint: Wang J, “ComBat-met: Adjusting Batch Effects in DNA Methylation Data.” bioRxiv, 2024. doi: 10.1101/2024.08.13.607838
1) Performed denoising, taxonomic classification, diversity estimation, and differential abundance analysis to characterize the respiratory microbiome and mycobiome of COPD patients, linking microbial changes to host immune responses.
2) Utilized linear mixed models to identify differentially expressed transcripts, proteins, metabolites, and lipids in COPD patients.
3) Performed weighted gene-coexpression network analysis (WGCNA) of BAL fluid macrophages to elucidate mechanisms of surfactant dysregulation.
4) Employed elastic net, support vector machines, and neural networks to predict COPD status and severity from molecular profiling data.
- Keywords: Microbiome, Transcriptomics, Proteomics, Metabolomics, Statistics, Machine Learning
- Publication: Hristova et al., “Multiomics Links Global Surfactant Dysregulation with Airflow Obstruction and Emphysema in COPD.” ERJ Open Res, 9(3): 00378-2022. doi: 10.1183/23120541.00378-2022
1) Developed PepSetTest, a peptide-centric method as a Bioconductor R package, to infer differentially expressed proteins based on coordinated changes of peptides.
2) Validated PepSetTest and identified protein signatures of tamoxifen-resistant breast cancer using data from TCGA/CPTAC.
- Keywords: Proteomics, Statistics, Software Development
- GitHub: https://github.com/JmWangBio/PepSetTest
- Publication: Wang J., Novick S., “Peptide Set Test: a Peptide-Centric Strategy to Infer Differentially Expressed Proteins.” Bioinformatics, 2024, 40(5): btae270. doi: 10.1093/bioinformatics/btae270
Developed unsteady-state parsimonious flux balance analysis, leveraging Gurobi optimization, to infer metabolic fluxes in cancer cells influenced by cancer-associated fibroblasts.
- Keywords: Metabolomics, Optimization, Software Development
- GitHub: https://github.com/FinleyLabUSC/CRC-Cell-Constraint-Based-Metabolic-Model
- Publication: Wang J., “Elucidating Tumor-Stromal Metabolic Crosstalk in Colorectal Cancer through Integration of Constraint-Based Models and LC-MS Metabolomics.” Metab Eng, 2022, 69: 175-187. doi: 10.1016/j.ymben.2021.11.006
1) Applied machine learning and artificial intelligence to model temporal and spatial patterns in organoid formation, enabling the prediction of cellular behavior.
2) Directed the analysis of bulk RNA-seq data to validate the intended target effects and identify potential off-target effects of a synthetic circuit in CHO cells.
3) Led the analysis of single-cell RNA-seq data to nominate candidate driver genes of hepatic development.
- Keywords: Transcriptomics, Single-Cell Transcriptomics, Machine Learning
- Publication: Caliendo et al., “Customizable Gene Sensing and Response without Altering Endogenous Coding Sequences.” Nat Chem Biol, 2024. doi: 10.1038/s41589-024-01733-y